With an all-new design that looks great on macOS Big Sur, Xcode 12 has customizable font sizes for the navigator, streamlined code completion, and new document tabs. Xcode 12 builds Universal apps by default to support Mac with Apple Silicon, often without changing a single line of code.
BioEdit is not available for Mac but there are plenty of alternatives that runs on macOS with similar functionality. The most popular Mac alternative is UGENE, which is both free and Open Source. If that doesn't suit you, our users have ranked 27 alternatives to BioEdit and 18 are available for Mac so hopefully you can find a suitable replacement. BioEdit, a sequence alignment. 95/98/NT/2000/XP Finch TV, freely available, and freely redistributable chromatogram viewer for both Window and Mac OS. For DNA sequence assembly and analysis Sequence Scanner Software v1.0, a free software enables you to view, edit, print and export sequence data generated using the Applied Biosystems. BioEdit No ClustalW Rudimentary, can read PHYLIP. Free, GPL: No Mac OS, Windows Official website: Discovery Studio Yes Align123, ClustalW, S-ALIGN UPGMA, NJ, with bootstrap and best tree. Software is package of 7 interactive visual tools for multiple sequence alignments. Major focus is manipulating large alignments. Furthermore, users can customize menu shortcuts for editor window, import compatible formats directly from the clipboard, or view and manipulate up to 20000 sequences. To summarize, despite not being continued by the developer, and having an outdated look, BioEdit remains an important free alternative to more expensive utilities. BioEdit is free and provides many basic functions for protein and nucleic sequence editing, alignment, manipulation and analysis. The software can still be downloaded but it is no longer maintained.
Designed for macOS Big Sur.
Xcode 12 looks great on macOS Big Sur, with a navigator sidebar that goes to the top of the window and clear new toolbar buttons. The navigator defaults to a larger font that’s easier to read, while giving you multiple size choices. New document tabs make it easy to create a working set of files within your workspace.
Document tabs.
The new tab model lets you open a new tab with a double-click, or track the selected file as you click around the navigator. You can re-arrange the document tabs to create a working set of files for your current task, and configure how content is shown within each tab. The navigator tracks the open files within your tabs using strong selection.
Navigator font sizes.
The navigator now tracks the system setting for “Sidebar icon size” used in Finder and Mail. You can also choose a unique font size just for Xcode within Preferences, including the traditional dense information presentation, and up to large fonts and icon targets.
Code completion streamlined.
A new completion UI presents only the information you need, taking up less screen space as you type. And completions are presented much faster, so you can keep coding at maximum speed.
Redesigned organizer.
An all-new design groups all critical information about each of your apps together in one place. Choose any app from any of your teams, then quickly navigate to inspect crash logs, energy reports, and performance metrics, such as battery consumption and launch time of your apps when used by customers.
SwiftUI
SwiftUI offers new features, improved performance, and the power to do even more, all while maintaining a stable API that makes it easy to bring your existing SwiftUI code forward into Xcode 12. A brand new life cycle management API for apps built with SwiftUI lets you write your entire app in SwiftUI and share even more code across all Apple platforms. And a new widget platform built on SwiftUI lets you build widgets that work great on iPad, iPhone, and Mac. Your SwiftUI views can now be shared with other developers, and appear as first-class controls in the Xcode library. And your existing SwiftUI code continues to work, while providing faster performance, better diagnostics, and access to new controls.
Universal app ready.
Xcode 12 is built as a Universal app that runs 100% natively on Intel-based CPUs and Apple Silicon for great performance and a snappy interface.* It also includes a unified macOS SDK that includes all the frameworks, compilers, debuggers, and other tools you need to build apps that run natively on Apple Silicon and the Intel x86_64 CPU.
Updated automatically
When you open your project in Xcode 12, your app is automatically updated to produce release builds and archives as Universal apps. When you build your app, Xcode produces one binary “slice” for Apple Silicon and one for the Intel x86_64 CPU, then wraps them together as a single app bundle to share or submit to the Mac App Store. You can test this at any time by selecting “Any Mac” as the target in the toolbar.
Test multiple architectures.
On the new Mac with Apple Silicon, you can run and debug apps running on either the native architecture or on Intel virtualization by selecting “My Mac (Rosetta)” in the toolbar.
Multiplatform template
New multiplatform app templates set up new projects to easily share code among iOS, iPadOS, and macOS using SwiftUI and the new lifecycle APIs. The project structure encourages sharing code across all platforms, while creating special custom experiences for each platform where it makes sense for your app.
Improved auto-indentation
Swift code is auto-formatted as you type to make common Swift code patterns look much better, including special support for the “guard” command.
StoreKit testing
New tools in Xcode let you create StoreKit files that describe the various subscription and in-app purchase products your app can offer, and create test scenarios to make sure everything works great for your customers — all locally testable on your Mac.
Bioedit Software Free For Mac Download
Get started.
Download Xcode 12 and use these resources to build apps for all Apple platforms.
This weeks’ lab will be held in Room B8218 (the Biology PC lab).Raw sequence files will be edited this week, and the edited sequence files will be analyzed next week.
Bioedit Software Free For Mac Os
Editing (and most of the analysis) will be done using BioEdit, a freeware sequence analysis program developed by Tom Hall at North Carolina State University.
Logging on
Each group should log on to a PC using the class ID bisc431 and the password pseud
Do not save your individual settings.
Sequence files
There are 4 disks containing sequence files.Each group should choose one of the sequence files on the disk, and copy it from drive A to the desktop.Also copy the file pstblue1vector.txt to the desktop.Return the disk as soon as this is done.
Opening BioEdit
Click on Start, Programs, and Bioedit.(You may have to scroll down the program list to find it.)
Opening files
Click on File menu, Open.Look in Desktop, files of type All files.Click on the sequence file you transferred to open it.(BioEdit should recognise it as an ABI Autosequencer Trace file and open it as a chromatogram.)
Editing sequence
Adjust the size of the chromatogram trace with the Horizontal scale and Vertical scale bars to the top left of the image.You should be able to clearly see the peaks of the trace.
Click on the view menu, and check editable sequence.This creates a duplicate sequence that can be edited without changing the original sequence.
The computer will already have called most of the bases from the peaks presentHowever, you will still see some “N”s in the sequence where the computer cannot make a call.By looking at the trace below the N, you should be able to make a visual judgement about which base should be present instead of N.(Each line in the trace is colour-coded to match the colour that one of the 4 bases is displayed in.)Select the N and replace it by typing in the appropriate base.(Note that sequences after 400-500 bases become increasingly unreliable, and are not worth spending much time on.)
Saving the edited file
Click on the File menu, Export as text.Save the edited file to the desktop, (or preferably, your own disk or network account.)
Preparing the Reverse Complement
The sequence present in the original file is the sequence of the newly synthesized strand.To get the sequence of the original template strand, the Reverse Complement must be prepared.
Click on the view menu (for the original unedited file), and check Reverse Complement.Note that this is also displayed in a 5'-3' direction, so the sequence complementary to the beginning of your original unedited forward sequence will be at the end of the reverse complement.Save the reverse complement as a text file under a different name.
Removing vector sequences
Bioedit Software For Windows 10
The sequences you are working with were prepared by the Davidson lab from DNA fragments cloned in the pSTBlue-1 plasmid vector.(See sequence analysis references for full map.)The clones were sequenced using either the T7 or SP6 promoter primers that flank the multiple cloning site in this vector.Because of this, the bases at the beginning of each sequence file you have are vector sequence, rather than cloned sequence.Since this may interfere with analysis of the sequence, these will have to be edited out.
To identify vector sequences, alignments will be prepared between your edited forward and reverse complement sequences and the sequence in the pstblue1vector.txt file.(This file contains the sequence of the multiple cloning site region of pSTBlue-1.)
Bioedit software, free download For Mac
Click on the File menu, New alignment.
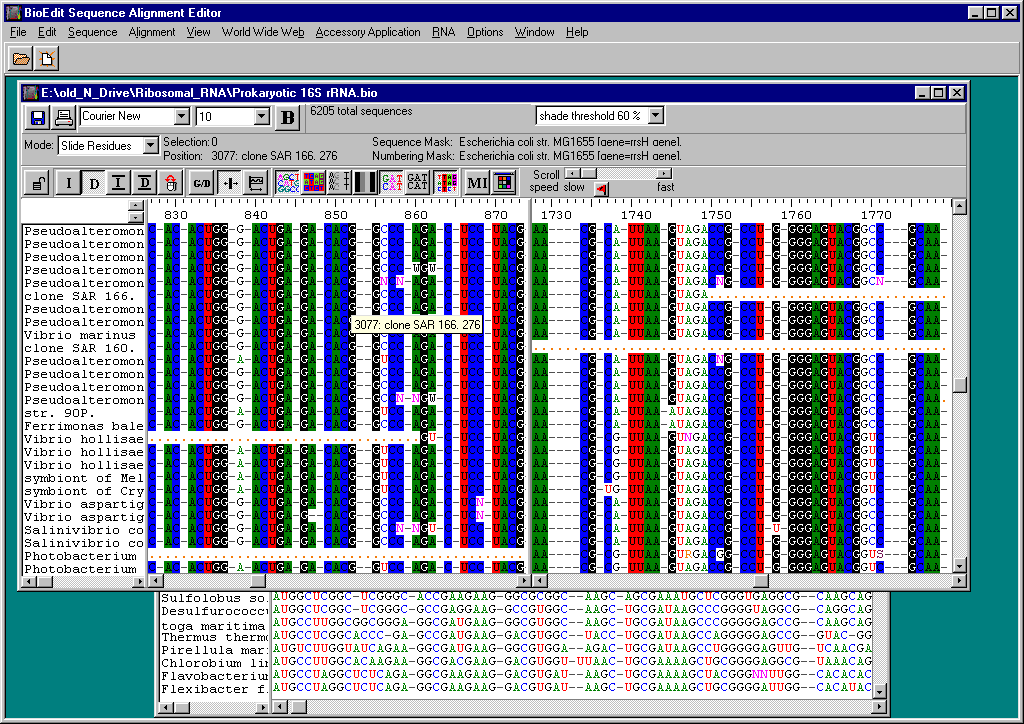
Click on the File menu, Import.Look in the Desktop (or wherever else you saved the edited sequence files), files of type All files.Click on the edited forward sequence file to open it.Repeat this process for the pstblue1vector.txt file.Select both files with the mouse by dragging it over the file names at the left.Click on Sequence menu, Pairwise alignment, Aligntwo sequence (allow ends to slide).If the vector sequence is on the same strand as the forward sequence, the vector should have a region of exact (or almost exact) homology with the beginning of the forward sequence.If this does not occur, repeat the process with the reverse complement sequence file (in a New alignment).If the vector sequence given is the opposite strand to the forward sequence, then there should be a region of (almost) exact homology with the end of the reverse complement.
Identify the region of vector sequences.(These should show an almost exact match to the forward or reverse sequence.If the alignment starts showing gaps and/or mismatches, the end of the vector sequence has been reached.)Return to your edited forward sequence file, delete the vector sequences, and save for next week.
